Virtual Coupling
When the chemical shift difference between two J coupled nuclei is of the same order as the coupling constant, second order spectra are obtained. See
this and
this. One, often unrecognized, second order effect is virtual coupling which is often misinterpreted as first order weak coupling. In a three-spin system, virtual coupling occurs when the observed nucleus appears to be coupled to both of the other two nuclei even though it is only coupled to one of them. This arises in AA'X and ABX spin systems when X (the observed nucleus) is coupled to only one of the other two strongly coupled spins. This is illustrated in the figure below.

The figure consists of simulations of X in an AA'X spin system as a function of JAA' with JAX set at 10 Hz and no coupling between A' and X. Clearly, the spectrum of X is affected by the coupling between A and A'. When JAA' = 0, a first order doublet is observed with a coupling constant of 10 Hz. As JAA' increases, complicated second order multiplets are observed. When JAA' = 50 Hz (or more) a "virtual triplet" with a coupling constant of 5 Hz is observed. This appears to be identical to a 1:2:1 triplet in a first order spectrum with a coupling constant of 1/2 JAX. It is however a second order spectrum and should not be misinterpreted as first order weak coupling. An example of this is illustrated in the figure below.
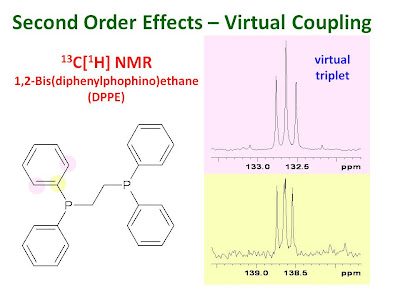
The figure shows the 13C NMR signals for the
ipso and
ortho aromatic carbons of 1,2-bis(diphenylphosphino)ethane (DPPE). These carbon atoms are coupled to the nearest phosphorus but not to the remote phosphorus. The two phosphorus atoms are strongly coupled to one another. The
ortho carbons appear as a "virtual triplet" and the
ipso carbons are second order multiplets.



Source:
University of Ottawa NMR Facility Blog



