Comparison of Broadband Decoupling Schemes
Many NMR measurements such as
HSQC or HMQC rely on
broadband X nucleus decoupling (X = 13C, 15N, 31P .... etc.).. Broadband decoupling schemes, using conventional rectangular pulses (e.g GARP) require fairly high power levels leading to undesired sample heating. They are also
limited in their effective decoupling bandwidth. Adiabatic decoupling schemes use
shaped adiabatic pulses and have become more and more common over the last couple of decades due, in large part, to the flexibility of modern NMR instruments to generate
shaped pulses. Adiabatic decoupling schemes (e.g. WURST) use much less power than those using conventional rectangular pulses. thereby reducing or eliminating problems associated with sample heating. Due to the lower power requirements and increased effectiveness over wider frequency ranges, adiabatic decoupling schemes are ideally suited for X nucleus decoupling at higher field strengths. The figure below shows 500 MHz 1H [31P] NMR spectra measured with
inverse gated decoupling for the P-CH3 methyl resonance of dimethyl methylphosphonate The single scan spectra were collected in a pseudo-2D fashion, as a function of the decoupler offset frequency from -256 ppm to +256 ppm from the 31P resonance frequency in 1 ppm steps. The acquisition time and recycle time for each FID were 2 sec and 4 sec, respectively. In the right-hand panel, broadband GARP decoupling was employed at a power of 1.22 W (60 µsec 90° pulses). In the left-hand panel, WURST decoupling was used at a peak power of 0.755 W (2 ms WURST pulses, bandwidth = 250 ppm).
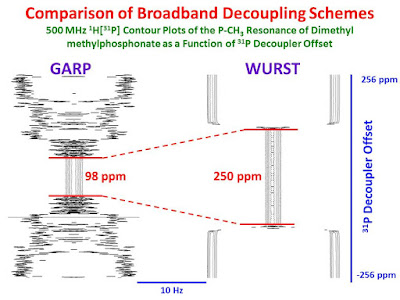
Clearly, the data collected with WURST decoupling, at lower power, have a much larger decoupling range (250 ppm) compared to the data collected with GARP decoupling (98 ppm). Furthermore, while the GARP data were collected, the sample temperature increased and had to be compensated for by the variable temperature unit. No such temperature increase was observed while collecting the WURST data. It is also interesting to note that, in the case of GARP decoupling,
distorted line shapes are observed just outside of the decoupling range, while for WURST decoupling, the spectra are fully coupled just outside of the decoupling range with a very sharp transition between being fully coupled and fully decoupled. For broadband decoupling, WURST is best!


Source:
University of Ottawa NMR Facility Blog



